Hi everyone,
I am trying to utilize a metaheuristic technique called SOS to do optimization inside Dynamo, and Python.
However, I can not release the output of calculation in Python inside Dynamo, as below:
It would be nice if Dynamo can release similar output as shown by utilizing Visual Studio Code, as below:
Here I upload the code if you want to test:
SOS Code.py (4.3 KB)
Feel free to reply if you have any idea about this.
Thanks
Instrad of using print(), use the syntax OUT = variable. The Python nodes in dynamo always function in that manner, so print function doesnt really get used in this context.
Hmm, I’ve put OUT statement, but it is located inside of “def” (line 72). Unfortunately still does not work, as below:
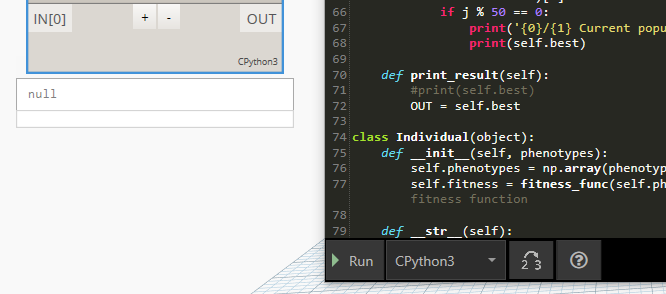
I think there is a difference for Python inside Dynamo
anyway, thank you for your reply

import numpy as np
class SOS(object):
def __init__(self,
l_bound,
u_bound,
population_size,
fitness_vector_size):
self.l_bound = l_bound
self.u_bound = u_bound
self.population_size = population_size
self.fitness_vector_size = fitness_vector_size
self.population = None
self.best = None
def float_rand(self, a, b, size=None):
return a + ((b - a) * np.random.random(size))
def generate_population(self):
population = self.float_rand(self.l_bound, self.u_bound, (self.population_size, self.fitness_vector_size))
self.population = np.array([Individual(p) for p in population])
self.best = sorted(self.population, key=lambda x: x.fitness)[0]
def mutualism(self, a_index):
b_index = np.random.permutation(np.delete(np.arange(self.population_size), a_index))[0]
b = self.population[b_index]
a = self.population[a_index]
bf1, bf2 = np.random.randint(1, 3, 2) # benefit factor1
array_rand = np.random.random(self.fitness_vector_size)
mutual = (a.phenotypes + b.phenotypes) / 2
new_a = a.phenotypes + (array_rand * (self.best.phenotypes - (mutual * bf1)))
new_b = b.phenotypes + (array_rand * (self.best.phenotypes - (mutual * bf2)))
new_a = Individual([self.u_bound if x > self.u_bound
else self.l_bound if x < self.l_bound else x for x in new_a])
new_b = Individual([self.u_bound if x > self.u_bound
else self.l_bound if x < self.l_bound else x for x in new_b])
self.population[a_index] = new_a if new_a.fitness < a.fitness else a
self.population[b_index] = new_b if new_b.fitness < b.fitness else b
def commensalism(self, a_index):
b_index = np.random.permutation(np.delete(np.arange(len(self.population)), a_index))[0]
b = self.population[b_index]
a = self.population[a_index]
array_rand = self.float_rand(-1, 1, self.fitness_vector_size)
new_a = a.phenotypes + (array_rand * (self.best.phenotypes - b.phenotypes))
new_a = Individual([self.u_bound if x > self.u_bound
else self.l_bound if x < self.l_bound
else x for x in new_a])
self.population[a_index] = new_a if new_a.fitness <= a.fitness else a
def parasitism(self, a_index):
parasite = np.array(self.population[a_index].phenotypes)
b_index = np.random.permutation(np.delete(np.arange(len(self.population)), a_index))[0]
b = self.population[b_index]
parasite[np.random.randint(0, self.fitness_vector_size)] = self.float_rand(self.l_bound, self.u_bound)
parasite = Individual(parasite)
self.population[b_index] = parasite if parasite.fitness <= b.fitness else b
def proceed(self, steps):
for j in range(1, steps + 1):
for i, val in enumerate(self.population):
self.mutualism(i)
self.commensalism(i)
self.parasitism(i)
self.best = sorted(self.population, key=lambda x: x.fitness)[0]
if j % 50 == 0:
print('{0}/{1} Current population:'.format(j, steps))
print(self.best)
def print_result(self):
print(self.best)
return self.best
class Individual(object):
def __init__(self, phenotypes):
self.phenotypes = np.array(phenotypes) # phenotype
self.fitness = fitness_func(self.phenotypes) #fitness function
def __str__(self):
return '{0} = {1}'.format(self.phenotypes, self.fitness)
def fitness_func(arg_vec):
s1 = (sum(arg_vec) - sum(x*x for x in arg_vec)) * sum(np.cos(x) for x in arg_vec)
s2 = 4 / (np.sqrt(np.abs(np.tan(sum(arg_vec))))) + int(sum(x*x for x in arg_vec))
return s1 / s2
interval = (-1, 1)
pop_size = 50 # candidate solutions
max_iter = 500 # iterations
dim = 5 # problem variables
sos = SOS(interval[0], interval[1], pop_size, dim)
sos.generate_population()
print('Initial population')
for ind in sorted(sos.population, key=lambda x: x.fitness):
print(ind)
max = sos.proceed(max_iter)
print('--------------------------')
OUT= np.asarray(sos.print_result() ).tolist()
Print statements in current builds should show up in the Dynamo console. Hit Shift+Up arrow to pull that open.
Hello,
the OUT variable must be declared (and set) in the global namespace
Thank you @RMohareb , It works.
Besides, thank you for your insights @jacob.small and @c.poupin





